IHC 2 APP: Ki-67 Nuclear Staining Analysis
The nuclear protein Ki-67 is an established cell proliferation marker in research and clinics. Its staining and evaluation in tissue sections is an integral part of cancer diagnosis and prognosis. Automated quantitative analysis of marker expression can provide a valuable supplement to visual evaluation. The IHC2 App is among the StrataQuest easiest-to-use analysis Apps and requires very little preparation time. This streamlined one-click analysis solution allows single cell detection, including the quantification of one additional marker within brightfield images.
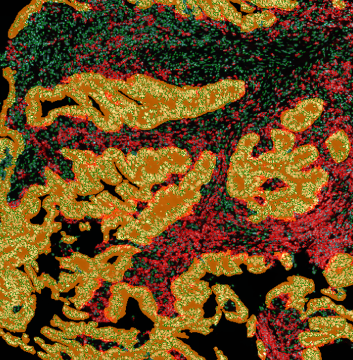
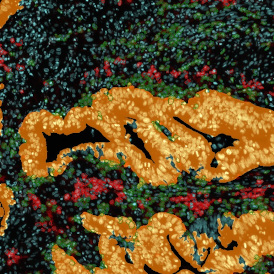
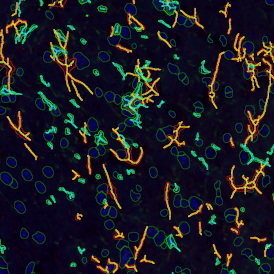
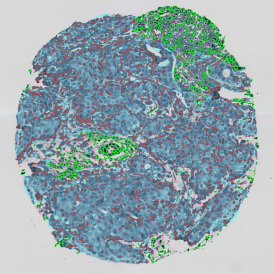
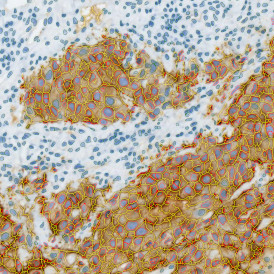
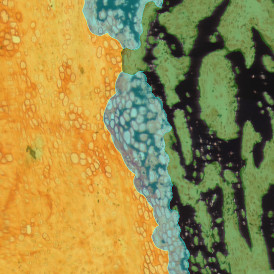
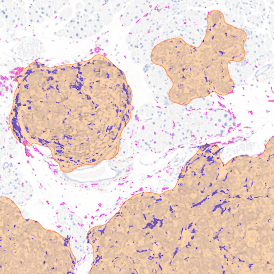
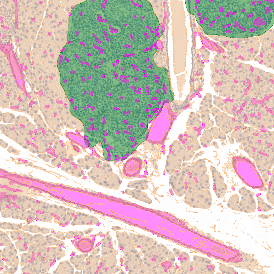
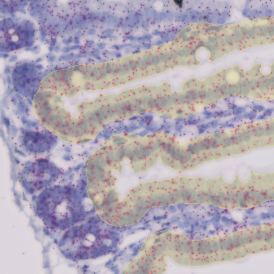
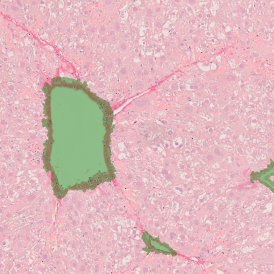
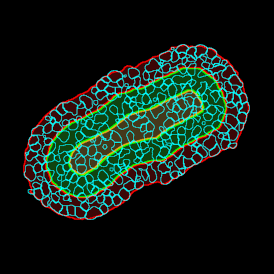
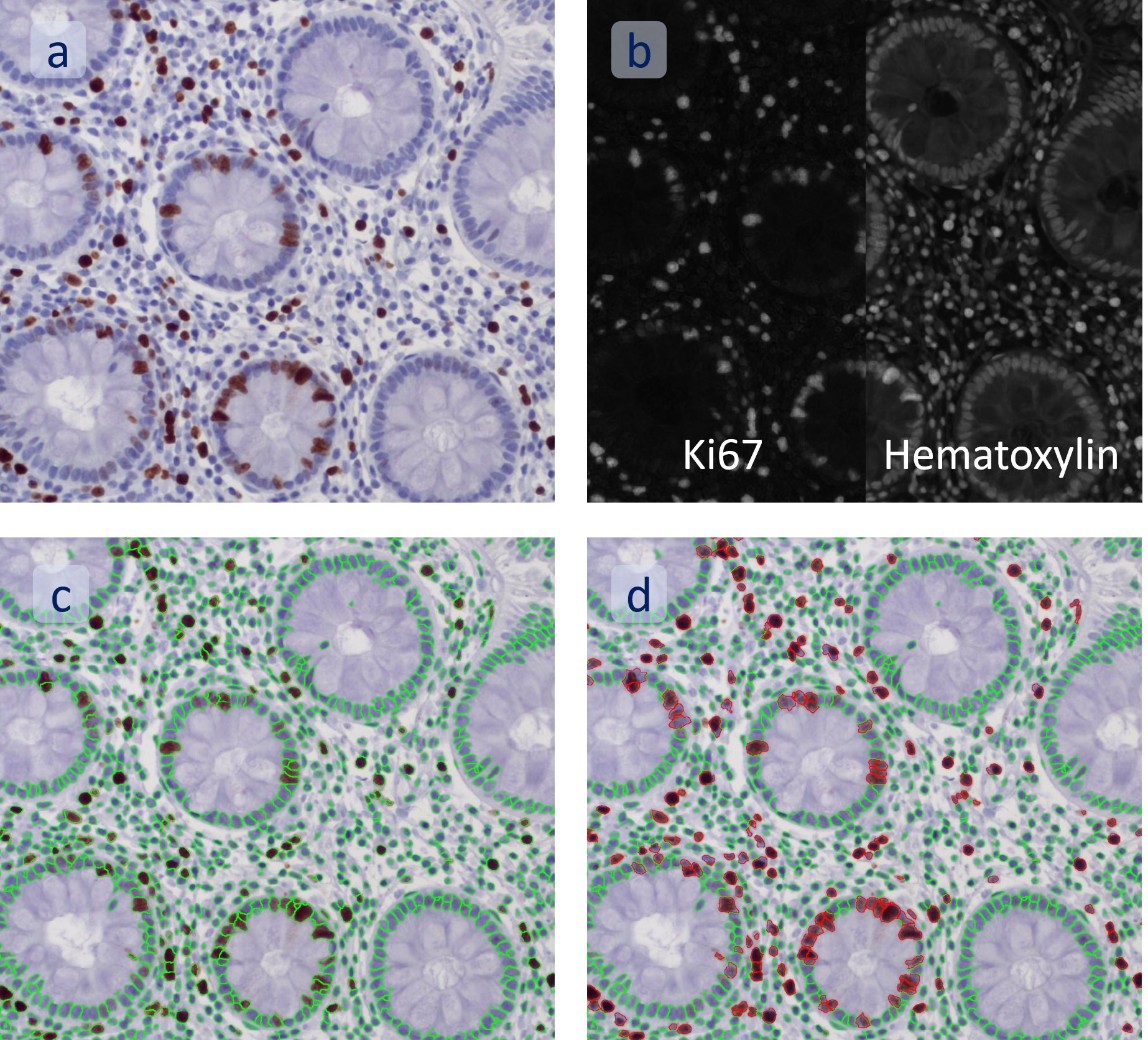
The aim of the following case study was to assess (i) the total number of cells as well as (ii) the number and % of Ki-67 positive cells. (a) shows colon tissue stained for hematoxylin (blue) to detect the nuclei and Ki-67 (brown).
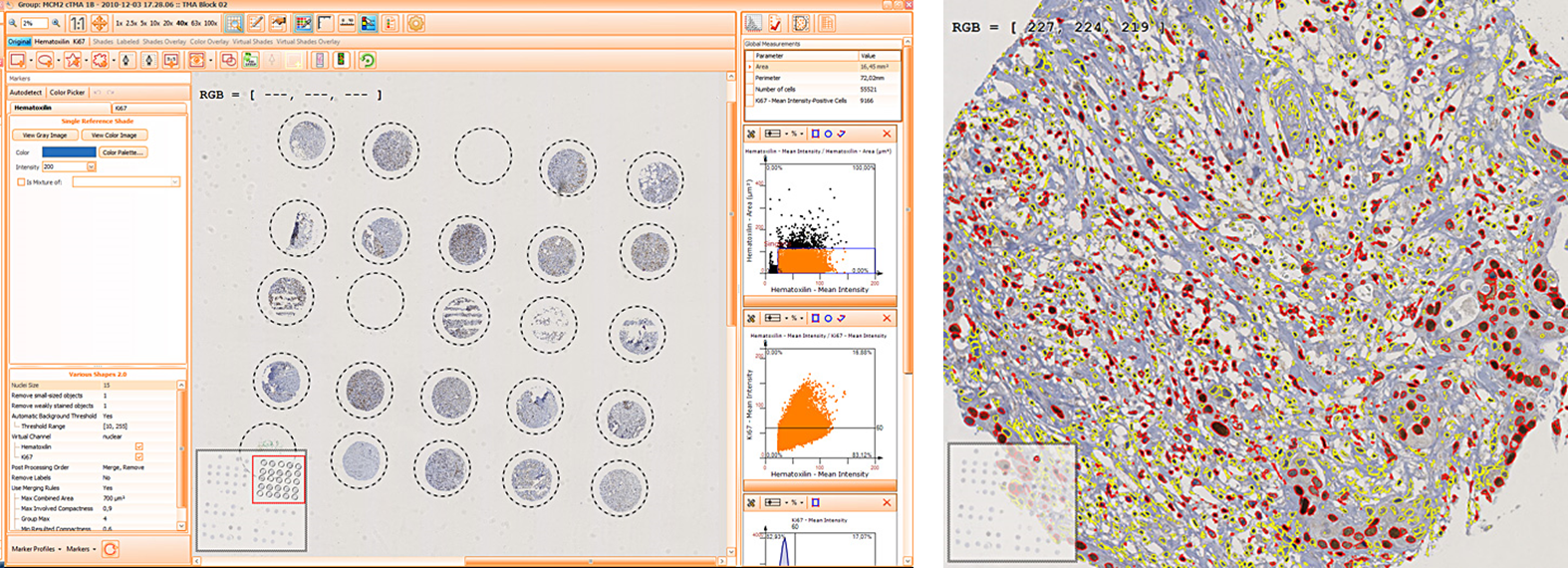
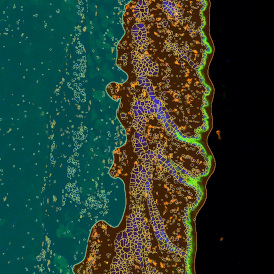
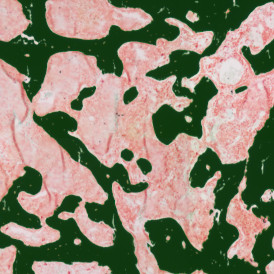
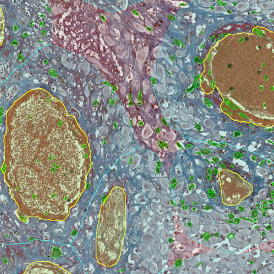
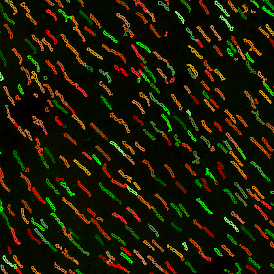
The first step of brightfield analysis is color separation. Using the color separation engine (b), the blue marker is automatically separated from the brown marker, as they are shown in grayscale images below.
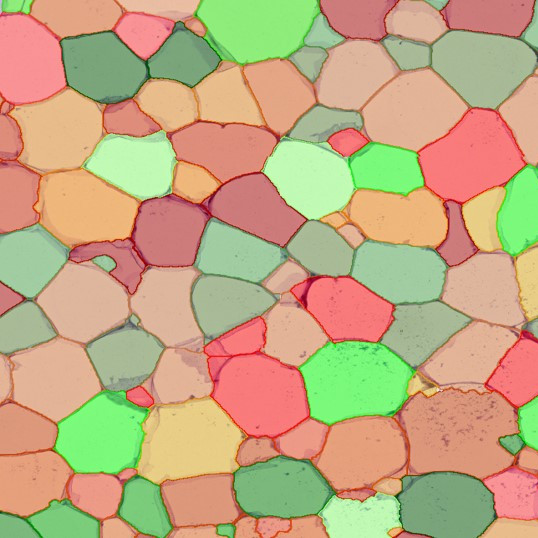
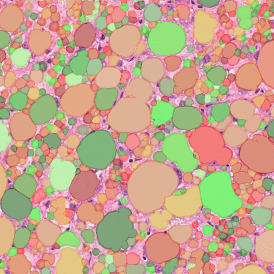
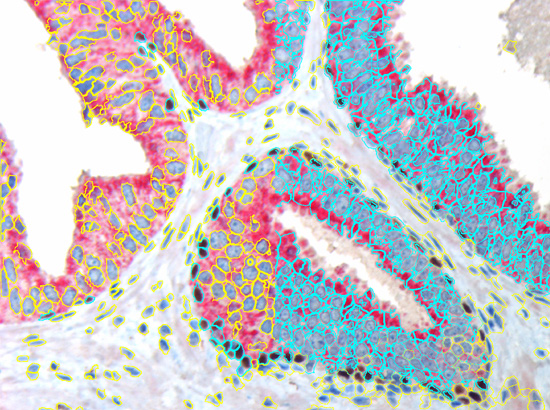
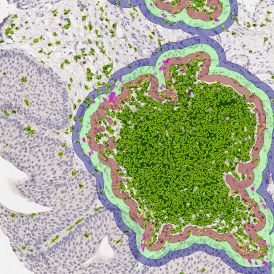
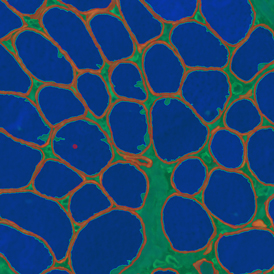
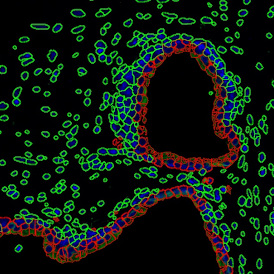
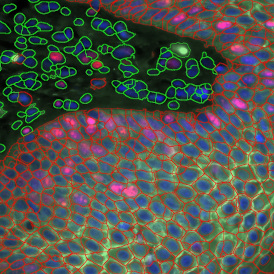
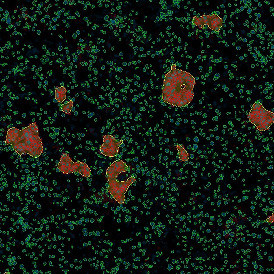
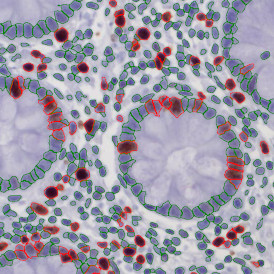
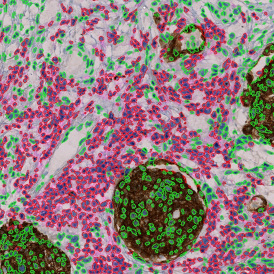
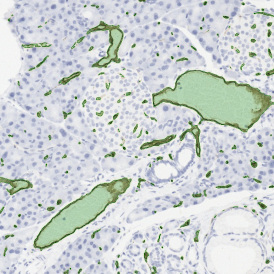
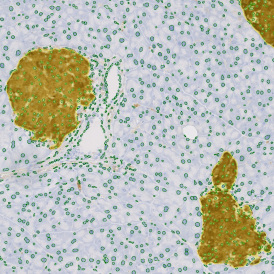
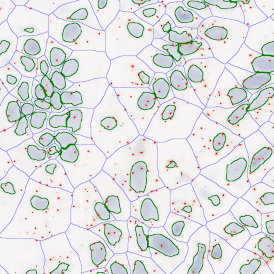
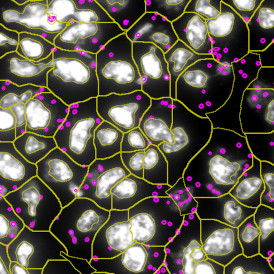
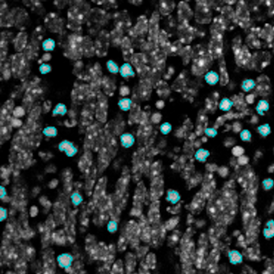
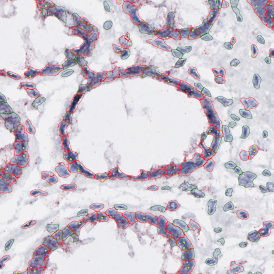
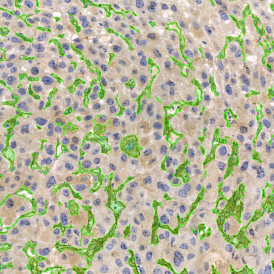
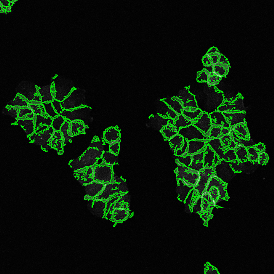
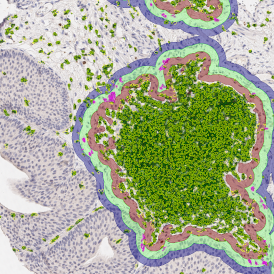
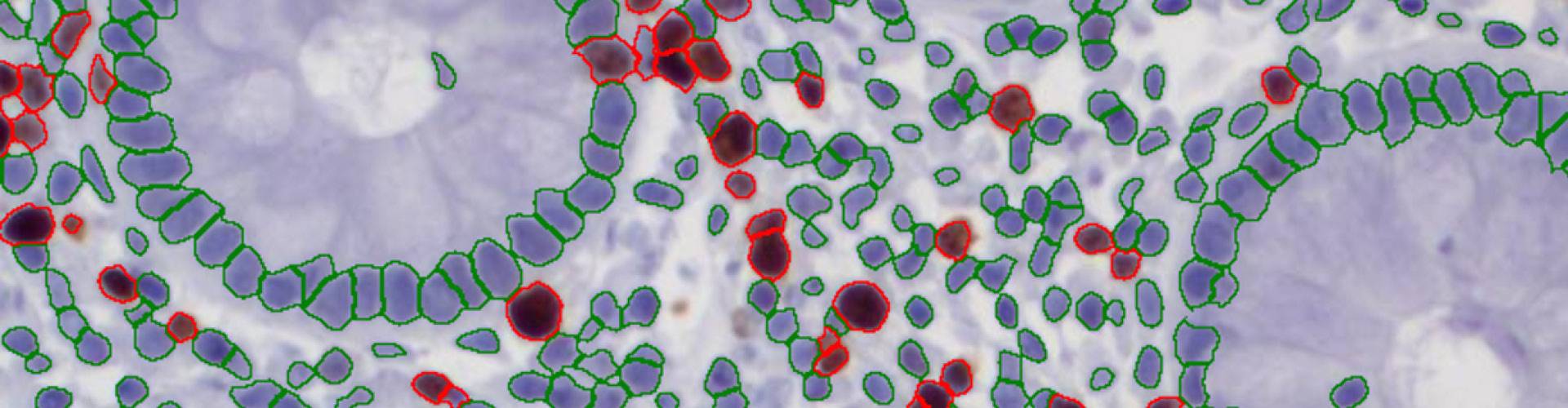
The second step is the automatic identification of nuclei based on hematoxylin and Ki-67 staining. Since Ki-67 is located in the nuclei and is also covering the hematoxylin staining there, the nuclei can be detected from both markers. Analysis masks provide visual control of the correctness of the nuclear segmentation – here the nuclei are outlined in green (c).
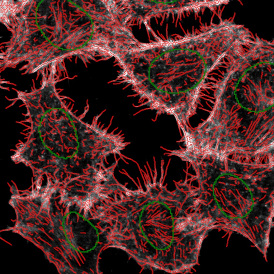
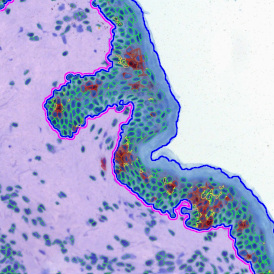
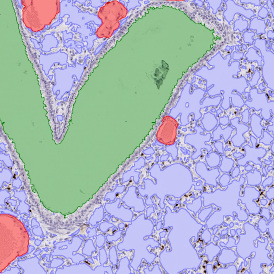
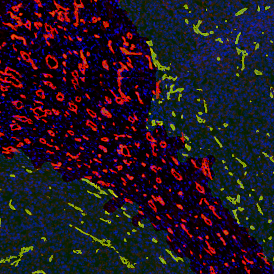
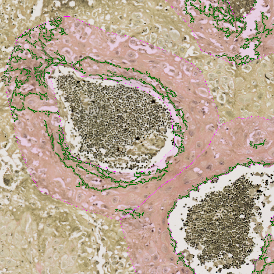
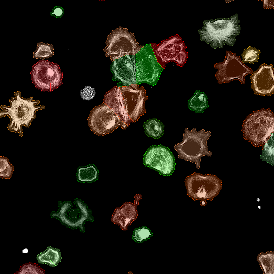
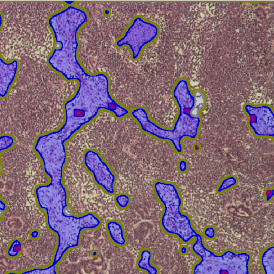
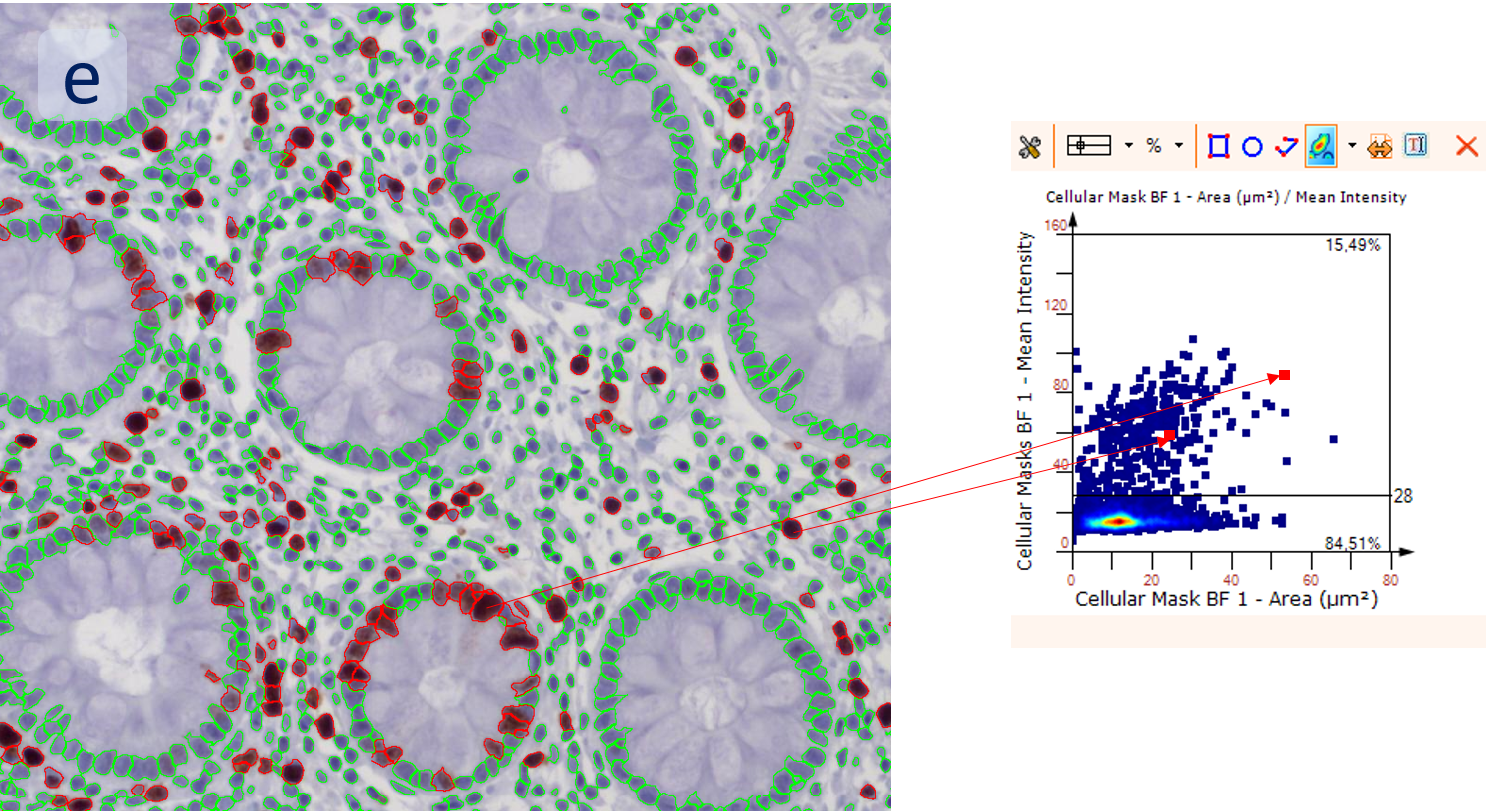
After setting a cut-off on the intensity of the Ki-67 marker in the corresponding scattergram, Ki-67 positive cells are shown with a red outline (d).
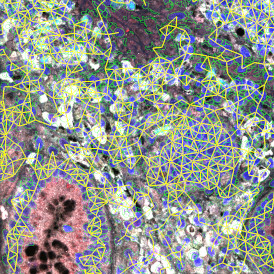
Backward connection allows to check cells of interest separately on the corresponding scattergram (e), for example, for checking a possible outlier. The scattergram on the example is plotted for mean Ki-67 intensity vs nuclei size. The results can be exported in common CSV or Excel formats.
Nuclear segmentation, as well as marker detection, is only one of the possibilities of the streamlined analysis solution available for StrataQuest. Get inspired by other analysis solutions in our StrataQuest App Center. Our team would also be glad to develop a personalized solution for you.


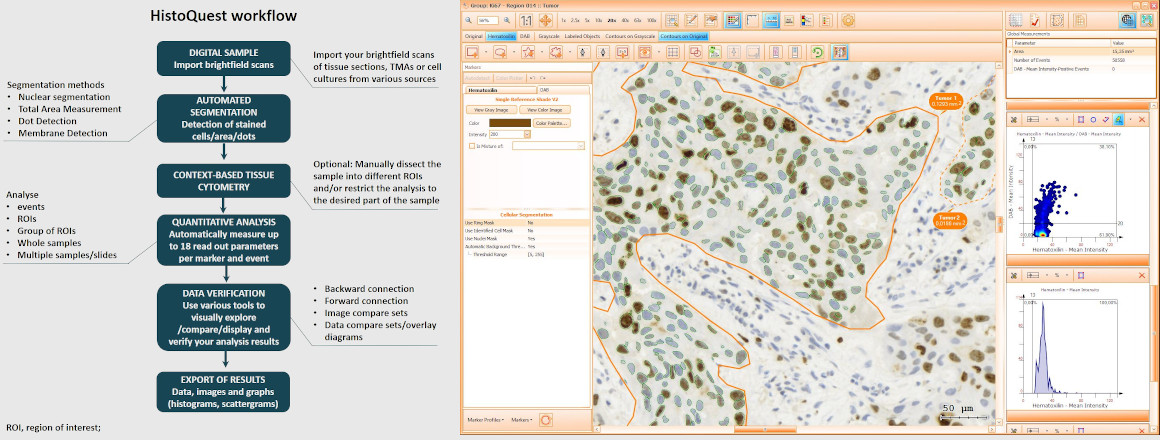
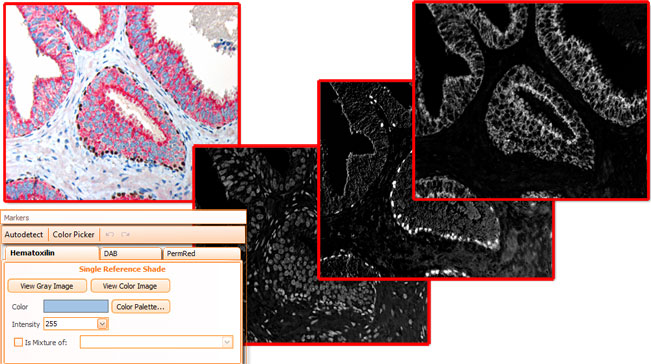 Brightfield images need to be separated into images according to their color components, HistoQuest provides two algorithms for color separation.
Brightfield images need to be separated into images according to their color components, HistoQuest provides two algorithms for color separation.