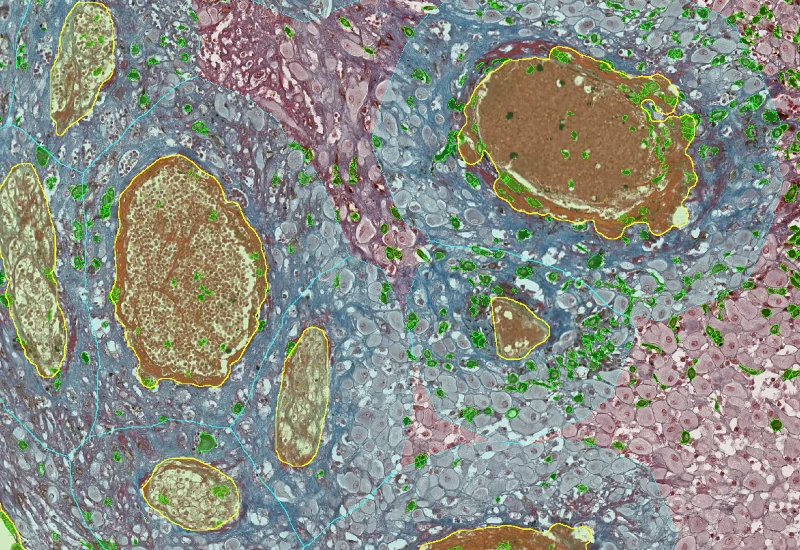
RNA Scope+
Detect nuclei and quantify two RNA scope dot markers per cell in nucleus and/or cytoplasm for CISH and SISH assays, measuring up to 20 morphometric and intensity parameters plus per-cell dot count, area, and intensity metrics.
metastructures
vascularization
CD31, blood vessels, diameter

The RNA Scope+ App provides detection of nuclei based on appropriate staining as well as dot detection per cell within nucleus and/or cytoplasm for two dot markers in CISH and SISH experiments. Each segmented cell compartment is measured for up to 20 intensity, statistic and morphometric parameters. Dot parameters are provided per cell and include count, mean intensity, total dot area, and sum of intensity as well as area and intensity lists for all single dots
Image courtesy of Melanie McCoy, The University of Western Australia
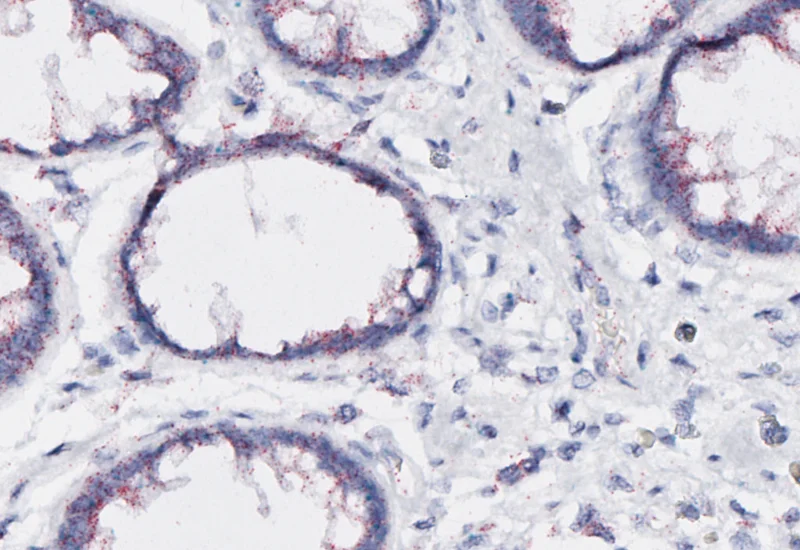
Original Image

Nuclei/cell detection

Red dot detection

Green dot detection
vascularization
Application Note
01 Jun, 2023
Evaluating the Distance of Tumor Cells from Blood Vessels
metastructures
Blog Post
17 May, 2023
An Intro to Deep Learning in Biomedical Imaging
We support the following file formats:
- TissueFAXS (aqproj)
- StrataFAXS II (vmic)
- PreciPoint (vmic, gtif)
- Generic BigTIFF Import
- Support for multipage BigTIFF files
- OME-TIFF
- JPEG, PNG, BMP, TIFF
- Zeiss (czi)
- Hamamatsu NanoZoomer (ndpi)
- Aperio (svs)
- Leica (scn)
- 3D HISTECH Pannoramic
- Mirax (mrxs)
- Olympus (vsi)
- More slide scanners to be added!
Related Apps
IHC Angio
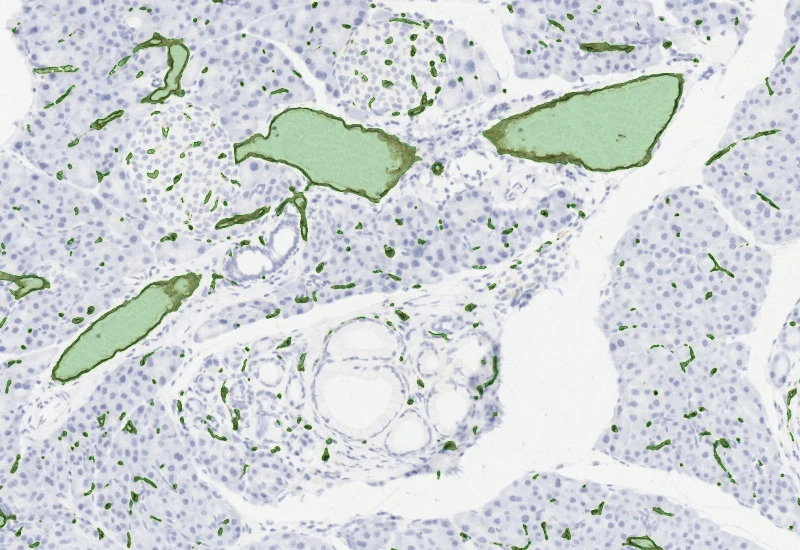
Detect blood vessels based on appropriate stains (e.g. CD31), measure vessel and lumen areas, and export vessel number, density, and areas of vessels, endothelium, and lumina.
blood vessels, tumor vascularization, tumor microenvironment, brightfield
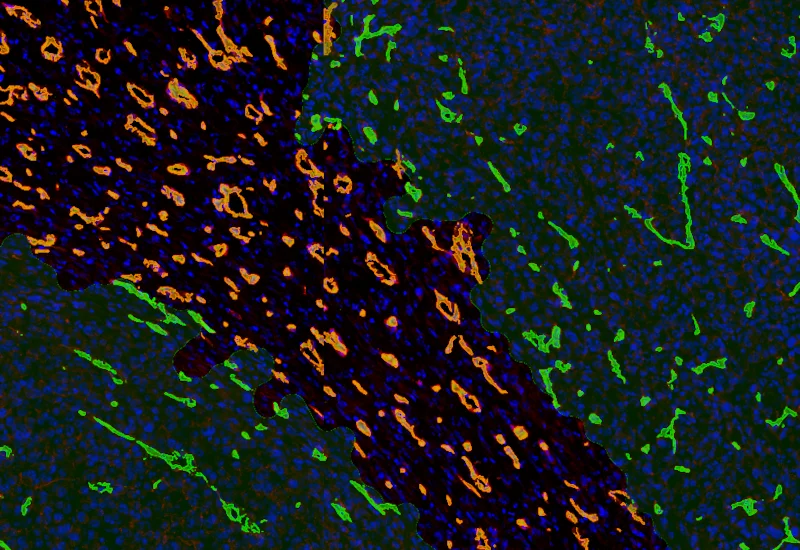
IF Tumor Vascularization
Segment tissue into tumor and stroma/healthy areas, detect CD31+ vessels, and quantify vessel number, area, density, and connectivity with configurable wall closing and distance linking.
vasculatization, cancer, stroma, tumor, blood vessels, CD31, spatial analysis, tumor microenvironment

Custom App development
Perfectly tailored image analysis solutions for your research.
You have a specific research question that needs to be answered? We offer custom development of image analysis pipelines for specific tasks, be it detection of cellular phenotypes or quantification of tissue structures. After discussing your goals with one of our experts, you will get a ready-to-use App and be a step closer to an impactful publication.