How immunofluorescence image analysis factors into NSCLC studies
It’s crucial to understand how immunofluorescence image analysis factors into NSCLC studies to uncover spatial biomarkers and guide research.
fluorescence
tumor microenvironment
oncology
immunology
Blog Post
How immunofluorescence image analysis factors into NSCLC studies
07 Apr, 2026
Non-small cell lung cancer (NSCLC) presents a significant research challenge due to its biological complexity, varied genetic drivers, and the intricate makeup of its surrounding microenvironment. Understanding how tumors behave within this landscape requires tools that go beyond genomic sequencing or conventional histology. Immunofluorescence image analysis addresses this need by enabling researchers to visualize and quantify protein expression within tissue samples, while preserving the spatial context of cells and structures. This dual capability is critical for investigating the mechanisms behind tumor progression, immune interaction, and therapeutic response.
The Role of Immunofluorescence Image Analysis in NSCLC Research
Detecting Biomarker Localization in Tumor Architecture
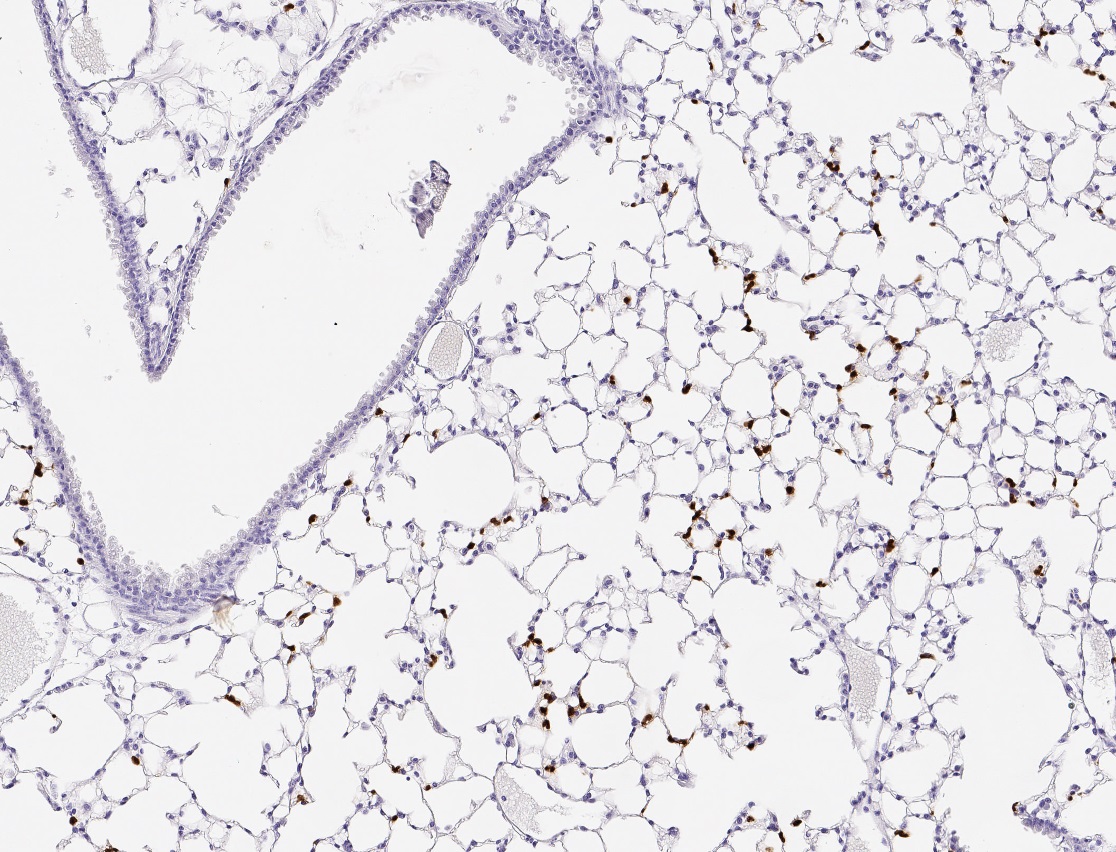
Image analysis of immunofluorescence stained NSCLC tissue samples allows for the precise localization of biomarkers like PD-L1, EGFR, and Ki-67 within the spatial context. It can reveal whether PD-L1 expression is limited to tumor cells or also present on immune cell populations. These distinctions are vital for assessing eligibility for immune checkpoint blockade therapy. Unlike homogenized assays, immunofluorescence image analysis investigates cellular phenotypes with their native tissue environment, allowing researchers to interpret biomarker relevance within spatial context.
Profiling the Tumor Immune Microenvironment
NSCLC tumors exhibit variable immune infiltration, which influences how patients respond to immunotherapy. Immunofluorescence enables the detection of multiple immune markers on a single tissue section. This helps researchers classify tumors into immune “hot” or “cold” types and investigate the spatial arrangement of various immune cells phenotypes around or within tumor nests, providing a more detailed view of immune evasion or activation.
Revealing Spatial Variability Within Tumors
Significant spatial variation in cellular phenotype and protein expression is often observed across a single NSCLC tumor. These differences are investigated using immunofluorescence image analysis, which highlights regions with distinct marker profiles. Identifying areas with elevated vimentin and reduced E-cadherin expression, for example, may indicate zones undergoing epithelial-to-mesenchymal transition, which is often linked to invasion and resistance.
Monitoring Longitudinal Changes in Treatment Response
Researchers assess tumor adaptation to therapy through immunofluorescence image analysis by comparing tissue samples taken before and after treatment. Shifts in marker expression, altered immune cell distribution, and newly emerging spatial patterns may signal the development of resistance. Such observations provide critical visual and quantitative evidence that can influence the refinement of therapeutic strategies in both clinical trials and laboratory models.
Linking Structure to Function in Multi-Omics Studies
When used alongside genomic, transcriptomic, or proteomic data, immunofluorescence image analysis anchors molecular findings in physical tissue structure. It helps correlate gene expression patterns with actual protein localization and cell behavior. This supports a more comprehensive understanding of NSCLC and enables researchers to develop more targeted hypotheses.
Implementing Immunofluorescence Image Analysis in NSCLC
Preparing the Sample for Multiplex Imaging
To begin the workflow, NSCLC tissue samples are sectioned and stained with panels of fluorescence tagged antibodies. These panels typically include tumor markers, immune-related proteins, and structural or lineage indicators, selected based on the biological questions under investigation.
Capturing and Processing High-Resolution Images
Following staining, slides are imaged using fluorescence or confocal whole slide scanners. Imaging systems record multiple channels corresponding to each marker.
Extracting Quantitative Insights from Spatial Data
Once the images are processed, cell segmentation algorithms identify individual nuclei and cell boundaries. Fluorescence intensity is then measured across defined regions, allowing researchers to quantify protein expression levels, identify co-localization patterns, and analyze spatial distributions across tumor zones or between cell types.
Key Applications of Immunofluorescence Image Analysis in NSCLC Research
Enhancing Diagnostic Confidence in Complex Cases
In certain NSCLC cases, especially those with poorly differentiated histology, immunofluorescence supports diagnosis through visualizing the expression of subtype-specific markers like TTF-1 or Napsin A. By preserving cell morphology and spatial relationships, the analysis helps pathologists reach more accurate conclusions even with limited tissue samples.
Identifying Predictive and Prognostic Biomarkers
Immunofluorescence image analysis contributes to biomarker validation using the quantitative measurement of protein expression within defined regions of the tumor microenvironment. A clear example is PD-L1 scoring, where its expression on tumor versus immune cells helps determine a patient’s eligibility for immunotherapy. Stratifying tumors this way improves trial design and guides the selection of treatments tailored to each patient's tumor biology.
Supporting Translational and Preclinical Research Models
NSCLC organoids, xenografts, and 3D culture systems are often evaluated using immunofluorescence image analysis to observe treatment effects and mimic patient tumor responses. This method allows for detailed phenotypic profiling. As a result, it enables researchers to assess biomarker expression, spatial organization, and cellular responses within preclinical models that closely resemble clinical tumors.
Elevating NSCLC Research Through Immunofluorescence Image Analysis
Immunofluorescence image analysis provides the spatial precision and molecular detail needed to better understand NSCLC at the cellular level. From mapping immune landscapes to tracking therapy response, it supports a wide range of research and clinical applications. For labs focused on high-resolution cellular analysis of fluorescence images, the StrataQuest image analysis platform from TissueGnostics offers powerful single-cell quantification in addition to neighbourhood analysis, tissue classification and many other tools, in a flexible, software-based environment. If you're exploring ways to advance your NSCLC research, connect with us and discover what’s possible with TissueGnostics.

