A New Cas9 mRNA‑Based Therapy Holds Promise to Treat Muscular Dystrophy
Quantitative confocal imaging helps researchers to confirm muscle protein restoration after Cas9 mRNA delivery via targeted lipid nanoparticles.
disease modeling
translational research
genomics
regenerative medicine
White Paper

A New Cas9 mRNA‑Based Therapy Holds Promise to Treat Muscular Dystrophy
13 May, 2026
Limb‑girdle muscular dystrophies (LGMDs) represent a heterogeneous group of inherited neuromuscular disorders characterized by progressive weakness of the shoulder and pelvic girdle muscles. Caused by mutations in more than 30 distinct muscle‑associated genes, LGMDs often lead to loss of independent mobility and significantly reduced quality of life (1). Despite advances in molecular diagnostics, disease‑modifying therapies remain limited for most LGMD subtypes, highlighting an urgent need for novel treatment strategies (2).
Among emerging approaches, CRISPR‑Cas9 genome editing has generated considerable excitement. By directly correcting disease‑causing mutations in skeletal and cardiac muscle, gene editing may lead to durable, potentially curative interventions.3 However, translating CRISPR‑based therapies into treatments for skeletal muscle disorders has faced substantial hurdles, particularly with respect to safe, efficient, and tissue‑specific delivery of gene‑editing tools.
Moving Beyond Viral Delivery: Selective Organ Targeting LNPs
Historically, adeno‑associated viral (AAV) vectors have been the dominant delivery platform for in vivo gene editing in muscles. While effective in preclinical models, AAV‑based approaches are restricted by high production costs, cargo size limits, immunogenicity, and the risk of persistent nuclease expression, which may increase off‑target editing (4-6).
A recent study addresses these limitations using selective organ targeting (SORT) lipid nanoparticles (LNPs) encapsulating Cas9 mRNA. LNPs provide a non‑viral, transient delivery system that allows repeated dosing and avoids long‑term nuclease expression. By fine‑tuning lipid composition, SORT LNPs can preferentially accumulate in specific tissues, including skeletal muscle (7). Researchers evaluated whether SORT LNPs carrying Cas9 cargo could efficiently edit muscle tissue in vivo and restore expression of Telethonin, a sarcomeric protein essential for muscle stability and force transmission.
Proof‑of‑Concept: Restoring Telethonin Expression in a Dystrophic Mouse Model
To demonstrate therapeutic potential, the team focused on limb‑girdle muscular dystrophy R7 (LGMDR7), caused by mutations in the TCAP gene encoding Telethonin. An engineered mouse model harboring a pathogenic microduplication in TCAP recapitulated the loss of Telethonin expression observed in patients.
The researchers compared SORT LNPs delivering Cas9 as either ribonucleoprotein (RNP) complexes or as mRNA. Notably, Cas9 mRNA‑loaded LNPs achieved markedly higher editing efficiency in skeletal muscle than their RNP counterparts. Following intramuscular injection, Cas9 mRNA SORT LNPs restored Telethonin expression to approximately 40 % of wild‑type levels in treated muscles (7).
Importantly, editing was not limited to the injection site. Quantifiable gene editing and protein restoration were also observed in adjacent muscle groups, indicating local distribution within muscle tissue. These findings suggest that Cas9 mRNA delivery via SORT LNPs can overcome some of the anatomical and physiological barriers that have historically limited non‑viral muscle targeting.
Quantitative Confocal Imaging Confirms Functional Protein Restoration
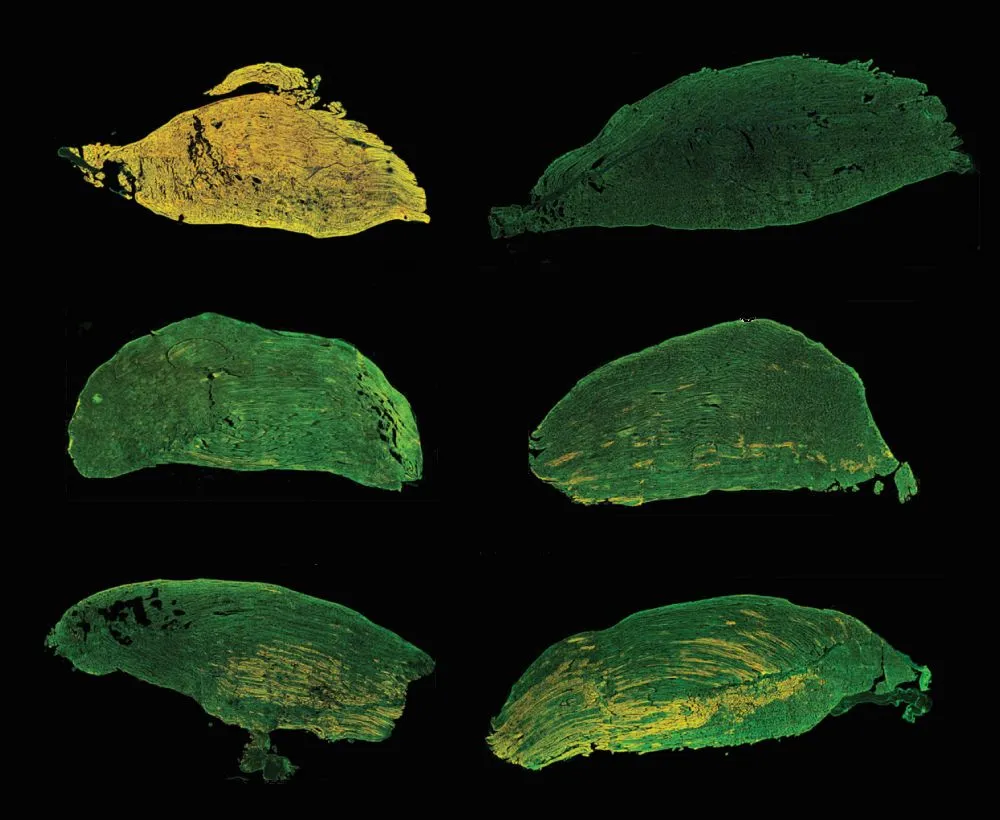
A key strength of the study lies in its rigorous validation of protein restoration at the tissue level. Whole‑slide confocal imaging of murine limb sections enabled direct comparison of muscle architecture and protein localization between wild‑type, untreated dystrophic, and Cas9 mRNA‑treated animals.
Using immunofluorescence staining for Telethonin and its binding partner Titin, the researchers confirmed that restored Telethonin localized correctly to the sarcomeric Z‑disc. This spatial accuracy is critical, as proper subcellular localization underpins mechanical stability and signaling within muscle fibers (8).
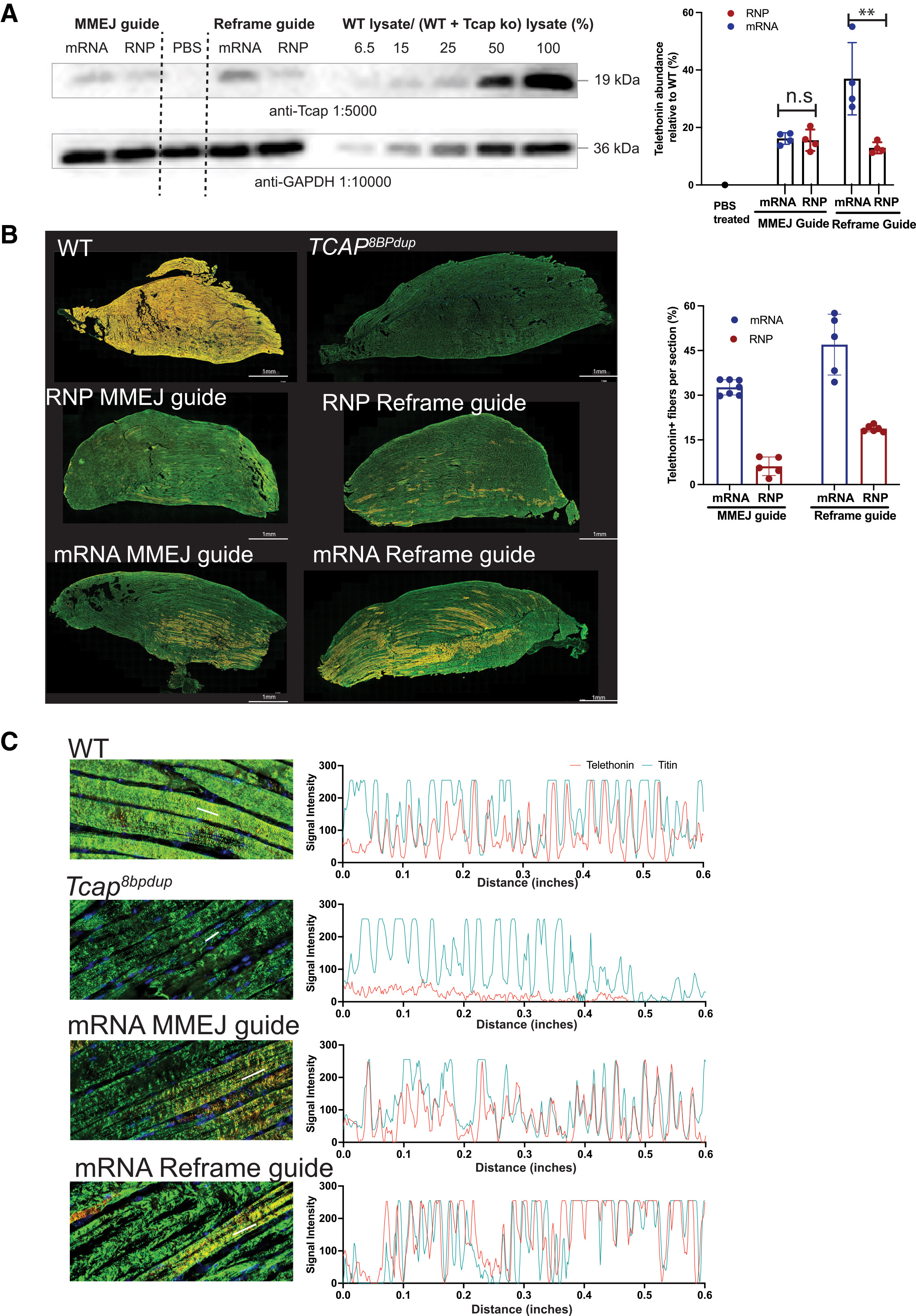
Source: Figure 4. Restoration of telethonin expression in myofibers upon Cas9 editing. Adapted from: Iyer S, Daman K, Sun Y et al, SORT LNPs encapsulating Cas9 mRNA achieve efficient editing in skeletal muscle in a dystrophic mouse model, Molecular Therapy, 2026.
High‑content, quantitative imaging was performed using TissueGnostics’s TissueFAXS SL Q slide scanner, which provided automated confocal whole‑slide acquisition across large tissue areas. The system’s motorized filter wheel with 10 positions and high‑resolution optics enabled reliable imaging of thick muscle sections stained for multiple markers, ensuring consistency across samples.
Subsequent analysis was carried out using StrataQuest image analysis software. This allowed automated tissue detection, segmentation of muscle fibers, and quantification of marker-positive cells. This combination of robust imaging hardware and flexible analysis software was essential for translating complex datasets into statistically meaningful biological conclusions.
Implications for Future Muscle Gene‑Editing Therapies
Beyond demonstrating proof‑of‑concept for LGMDR7, this study provides broader insights for the development of non‑viral gene‑editing therapies targeting skeletal muscle. The superiority of Cas9 mRNA over RNP delivery in large, multinucleated myofibers underscores the importance of cargo format, particle size, and intracellular kinetics for successful editing.
Immune profiling revealed distinct but manageable innate and adaptive immune responses to repeated LNP dosing, without loss of editing efficiency. This finding supports the feasibility of redosing strategies, an important advantage over viral vectors for chronic diseases such as muscular dystrophy.
Conclusion
Together, these findings demonstrate that Cas9 mRNA‑based gene editing delivered via SORT lipid nanoparticles can partially restore disease‑relevant protein expression in dystrophic skeletal muscle. Equally important, the study highlights the critical role of advanced quantitative imaging in validating potential therapeutic outcomes.
By using core facility–ready solutions such as our TissueFAXS SL Q slide scanner, which combines fast confocal imaging with high capacity of 120 slides, and StrataQuest analysis platform, the research team was able to confirm their hypothesis with high confidence and spatial precision. This technology combination enabled reproducible, whole‑tissue assessment of protein localization and expression levels across sample sets, supporting statistically robust conclusions.
As non‑viral genome editing continues to mature, integrated approaches that combine innovative delivery technologies with robust quantitative imaging will be essential for moving promising therapies from bench to bedside.
References
1. Bouchard, C., & Tremblay, J.‑P. (2023). Limb‑girdle muscular dystrophies: Classification and therapies. J. Clin. Med, 12, 4769.
2. Liewluck, T., and Milone, M. (2018). Untangling the complexity of limb-girdle muscular dystrophies. Muscle Nerve 58, 167–177.
3. Benarroch, L., et. al. (2024). The 2024 version of the gene table of neuromuscular disorders (nuclear genome). Neuromuscul. Disord. 34, 126–170.
4. Wang, D., Tai, P. W. L., & Gao, G. (2019). Adeno‑associated virus vector as a platform for gene therapy delivery. Nature Reviews Drug Discovery, 18, 358–378.
5. Padmaswari, M.H., et. al. (2023). Delivery challenges for CRISPR-Cas9 genome editing for Duchenne muscular dystrophy. Biophys. Rev. 4, 011307
6. Nelson, C.E., et al. (2019). Long-term evaluation of AAV-CRISPR genome editing for Duchenne muscular dystrophy. Nat. Med. 25, 427–432.
7. Iyer, S., et al. (2026). SORT LNPs encapsulating Cas9 mRNA achieve efficient editing in skeletal muscle in a dystrophic mouse model. Molecular Therapy, 34(7).
8. Zou, P., et al. (2006). Palindromic assembly of the giant muscle protein titin in the sarcomeric Z‑disk. Nature, 439, 229–233.

