IF Hi-plex 50
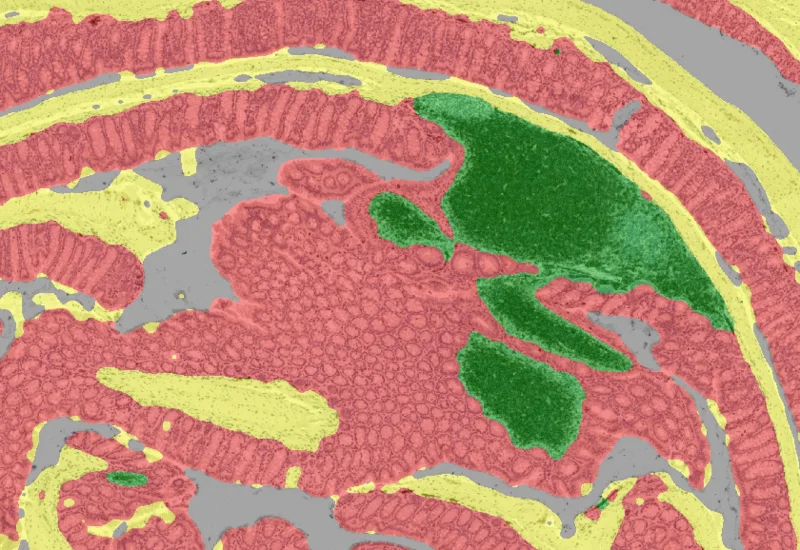
Combine and analyze up to 50 IF-stained images of the same tissue, define cellular phenotypes, segment cells into nucleus, perinuclear area, and/or cytoplasm.
single-cell analysis
tumor microenvironment
multiplex IF
immunophenotyping
multiplexing, immune cells, immune landscape, t cells, b cells, marcophages, monocytes, dentritic cells, MELC, skin, mouse, fluorescence

The IF Hi-Plex 50 App combines and analyses images of the same tissue section IF stained/acquired for up to 50 times with different markers. The APP allows to detect the cellular phenotypes of specific IF stained cell populations. It segments cells into nucleus, and/or perinuclear area and/or cytoplasm. Each segmented cell compartment is measured for up to 20 intensity, statistic and morphometric parameters which can be displayed in scattergrams and histograms and exported.

Original image

Nuclei detection

Marker detection

Hi-plex marker detection
immunophenotyping

Webinar
20 Jan, 2026
Decoding Metastatic Potential in Colorectal Cancer Using Tissue Cytometry
multiplex IF

White Paper
17 Oct, 2025
Integrative Multiomics Approach Unveils Systemic Dysfunction in Colorectal Cancer (CRC)
tumor microenvironment
Blog Post
07 Apr, 2026
How immunofluorescence image analysis factors into NSCLC studies
single-cell analysis

White Paper
30 Mar, 2026
Understanding NeuroCOVID-19: SARS-CoV-2 Disrupts Astrocyte Homeostatic Functions
We support the following file formats:
- TissueFAXS (aqproj)
- StrataFAXS II (vmic)
- PreciPoint (vmic, gtif)
- Generic BigTIFF Import
- Support for multipage BigTIFF files
- OME-TIFF
- JPEG, PNG, BMP, TIFF
- Zeiss (czi)
- Hamamatsu NanoZoomer (ndpi)
- Aperio (svs)
- Leica (scn)
- 3D HISTECH Pannoramic
- Mirax (mrxs)
- Olympus (vsi)
- More slide scanners to be added!
Related Apps
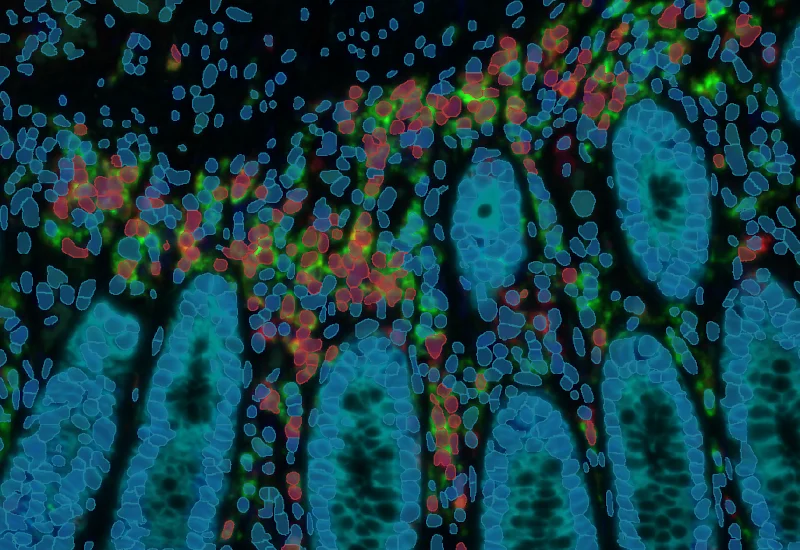
IF 4
Analyze single-cell co-expression of four IF markers, segment cells into nucleus, perinuclear area, and/or cytoplasm, and export up to 20 intensity, statistic, and morphometric parameters.
co-expression, phenotyping, fluorescence, epithelial cells, immune cells
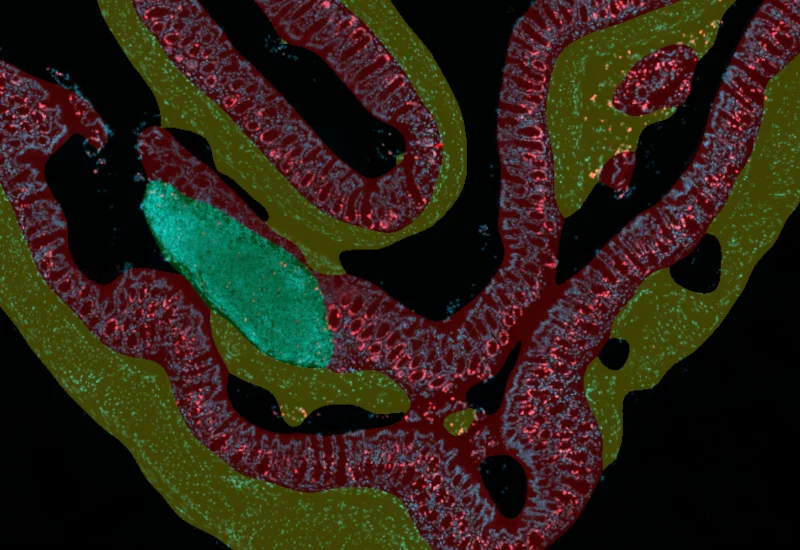
IF Swiss Roll
The IF Swiss Roll App segments tissue into subclasses (e.g., mucosa, follicles, connective tissue), detects nuclei, and identifies phenotypes via IF stains.
mouse, colon, fluorescence, immune cell follicles

Custom App development
Perfectly tailored image analysis solutions for your research.
You have a specific research question that needs to be answered? We offer custom development of image analysis pipelines for specific tasks, be it detection of cellular phenotypes or quantification of tissue structures. After discussing your goals with one of our experts, you will get a ready-to-use App and be a step closer to an impactful publication.