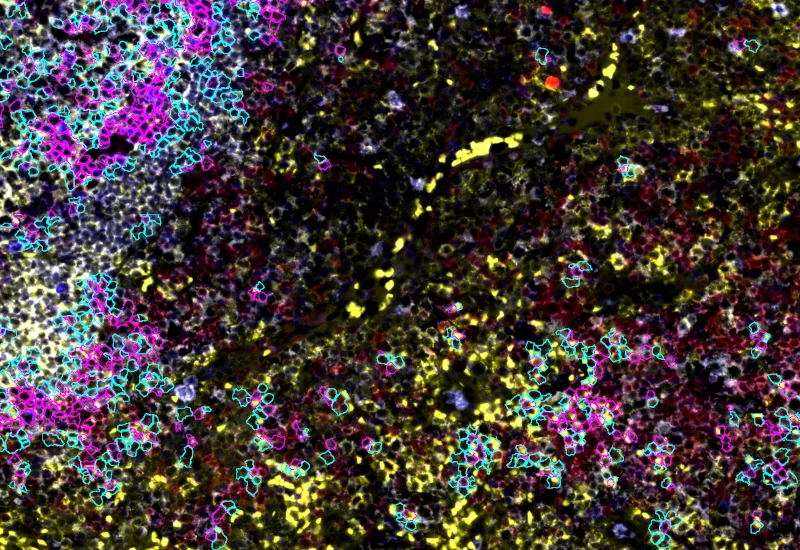
IF Cellular Contact
Determine cellular phenotypes in IF-stained samples, analyze direct cell–cell contacts, and quantify marker intensity, morphometric parameters, and number/% of contacting phenotypes.
single-cell analysis
tumor microenvironment
immunophenotyping
cell contacts, spatial interactions, phenotype interactions, B cells, T cells, lung, lymph node, COVID-19, SARS-Cov-2

The IF Cellular Contact App allows to determine the cellular phenotype of specific IF stained cell populations and establishes the cellular contacts to their neighboring cells (the number of markers is technically unlimited). If needed, the App provides a separation of nuclei in tissues with high cellular densities. The App outputs parameters such as staining intensity per marker and morphometric parameters for each segmented cell/cell compartment as well as number and percentage of cells of different phenotypes in direct contact.
Kaneko N, Boucau J, Kuo HH, et al. Temporal changes in T cell subsets and expansion of cytotoxic CD4+ T cells in the lungs in severe COVID-19. Clin Immunol. 2022;237:108991. doi:10.1016/j.clim.2022.108991
Image courtesy of Naoki Kaneko/Shiv Pillai (PI), Ragon Institute of MGH, MIT and Harvard, Boston, MA USA
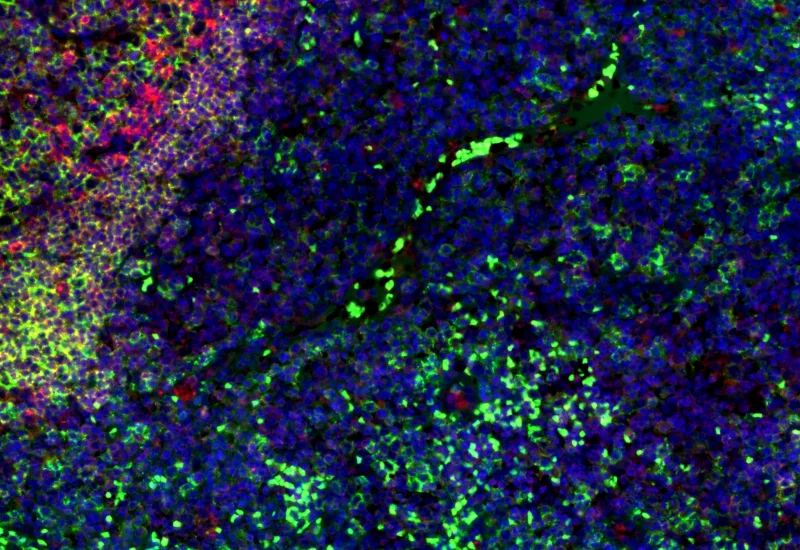
Original Image

Nuclei detection
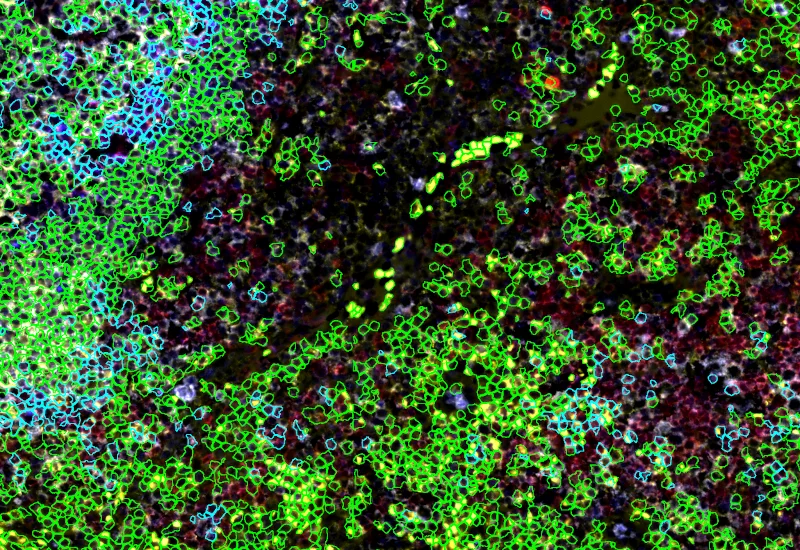
Phenotype 1 and Phenotype 2 detection
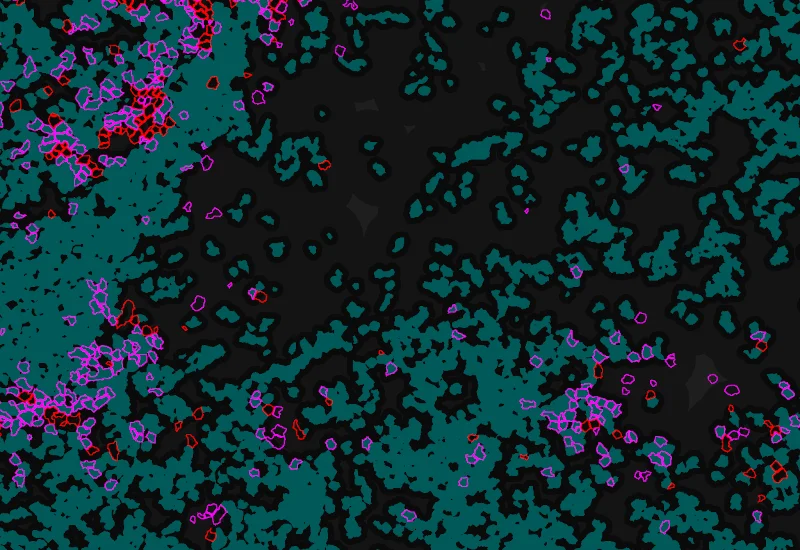

Phenotype 1 to 2 proximity 0-5μm
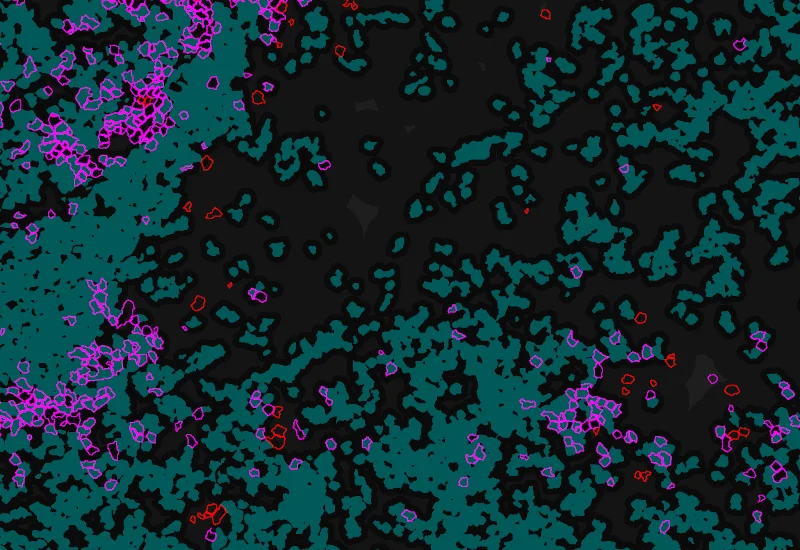

Phenotype 1 to 2 proximity 5-25μm

Phenotype 1 to 2 direct contact
immunophenotyping

Webinar
20 Jan, 2026
Decoding Metastatic Potential in Colorectal Cancer Using Tissue Cytometry
tumor microenvironment
Blog Post
07 Apr, 2026
How immunofluorescence image analysis factors into NSCLC studies
single-cell analysis

White Paper
30 Mar, 2026
Understanding NeuroCOVID-19: SARS-CoV-2 Disrupts Astrocyte Homeostatic Functions
We support the following file formats:
- TissueFAXS (aqproj)
- StrataFAXS II (vmic)
- PreciPoint (vmic, gtif)
- Generic BigTIFF Import
- Support for multipage BigTIFF files
- OME-TIFF
- JPEG, PNG, BMP, TIFF
- Zeiss (czi)
- Hamamatsu NanoZoomer (ndpi)
- Aperio (svs)
- Leica (scn)
- 3D HISTECH Pannoramic
- Mirax (mrxs)
- Olympus (vsi)
- More slide scanners to be added!
Related Apps
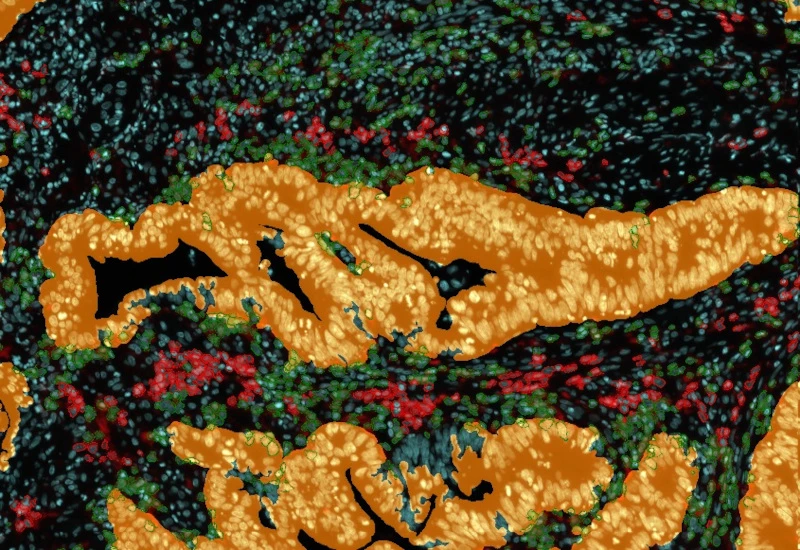
IF Immune status in situ
Characterize immune cell phenotypes relative to detected metastructures (e.g. tumors, glands), define distance ranges, measure cell-to-boundary distances inside/outside, and export up to 20 intensity, statistic, and morphometric parameters per cell compartment.
colon cancer, cytotoxic t cells, tumor microenvironment, PD1, CD8, spatial analysis, fluorescence
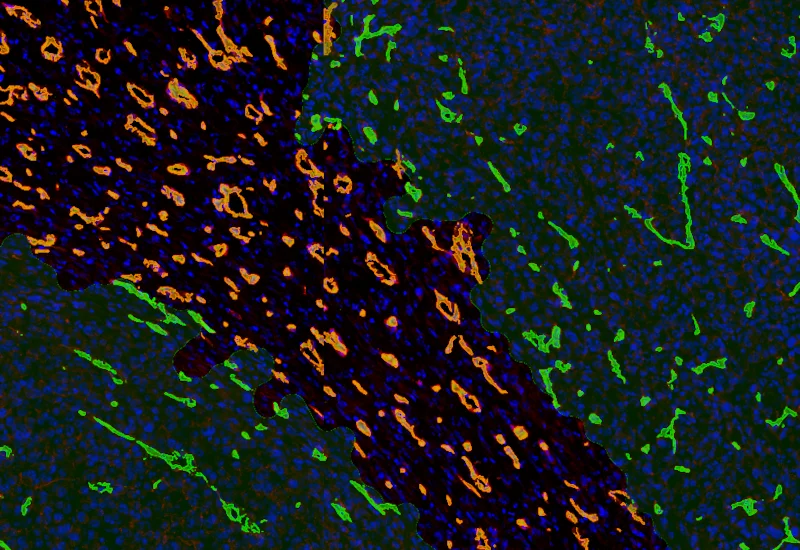
IF Tumor Vascularization
Segment tissue into tumor and stroma/healthy areas, detect CD31+ vessels, and quantify vessel number, area, density, and connectivity with configurable wall closing and distance linking.
vasculatization, cancer, stroma, tumor, blood vessels, CD31, spatial analysis, tumor microenvironment

IHC Immune Status in Situ
Segment tissue into tumor, stroma, and lymphocyte clusters using an AI classifier, detect hematoxylin-stained nuclei and immune phenotypes (e.g., CD45, CD3, CD20), and quantify spatial immune distribution.
bladder cancer, CD45, immune cells, tumor immune microenvironment, tertiary lymphoid structures, spatial analysis

Custom App development
Perfectly tailored image analysis solutions for your research.
You have a specific research question that needs to be answered? We offer custom development of image analysis pipelines for specific tasks, be it detection of cellular phenotypes or quantification of tissue structures. After discussing your goals with one of our experts, you will get a ready-to-use App and be a step closer to an impactful publication.

