IF Dots Colocalization
Detect dot-like signals (e.g., FISH, RNAscope) in multiple fluorescence channels and quantify dot number, size, and colocalization between markers.
FISH, cell culture, RNAScope, colocalization

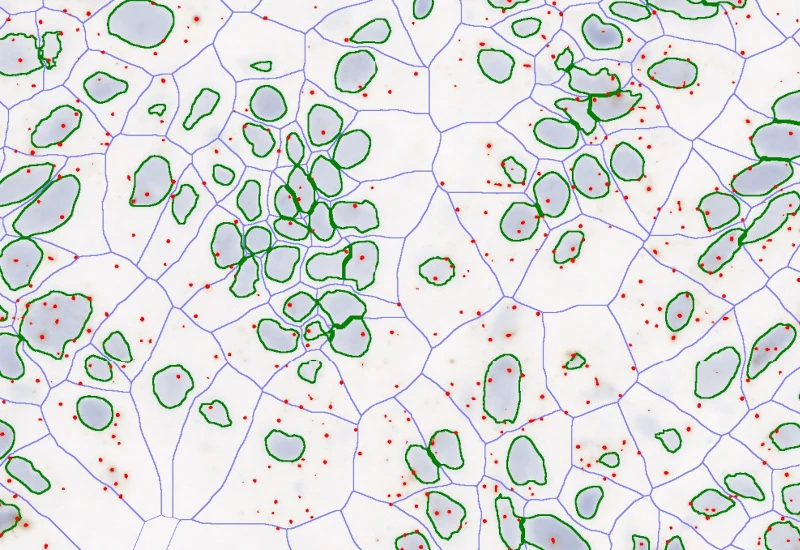
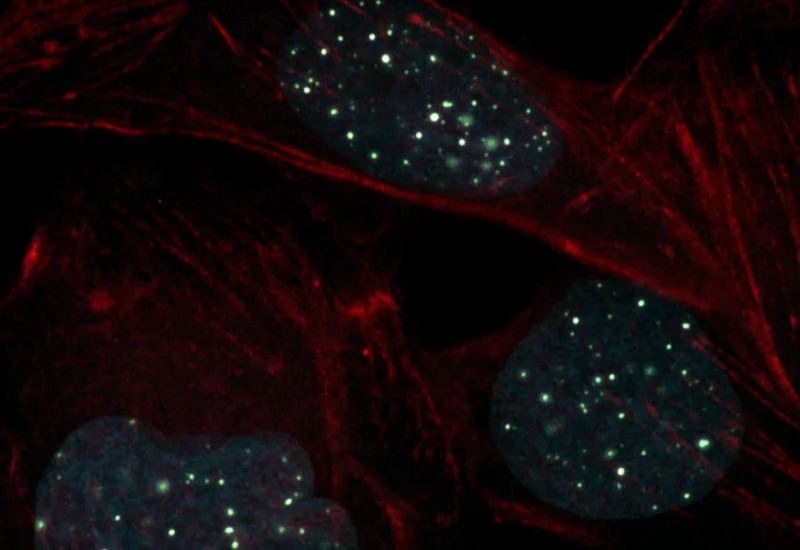
The IF Dots Colocalization App allows the detection of dot-like stainings (e.g., FISH, RNAscope) and the analysis of their colocalization. The output includes the number/area of detected dots as well as the number/area of colocalized dots.
Vina Putra, Nuclear Dynamics Group, CMRI, Westmead

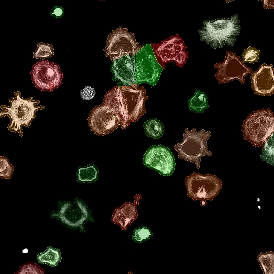
Original Image

Detection of green dots

Detection of pink dots

Identification of colocalized dots
Customer Publication
12 Oct, 2021
Tissue Cytometry in HIV Research
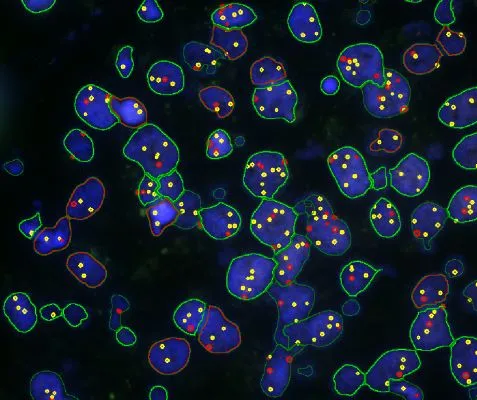
Application Note
14 Feb, 2023
Case Study: Analysis of FISH Using Tissue Cytometry
Blog Post
15 Feb, 2023
Applications of AI in Cell Segmentation
We support the following file formats:
- TissueFAXS (aqproj)
- StrataFAXS II (vmic)
- PreciPoint (vmic, gtif)
- Generic BigTIFF Import
- Support for multipage BigTIFF files
- OME-TIFF
- JPEG, PNG, BMP, TIFF
- Zeiss (czi)
- Hamamatsu NanoZoomer (ndpi)
- Aperio (svs)
- Leica (scn)
- 3D HISTECH Pannoramic
- Mirax (mrxs)
- Olympus (vsi)
- More slide scanners to be added!
Related Apps

IF Dots
Detect dots-stainings per cell in up to four markers, segment cellular compartments, measure up to 20 intensity, statistic, and morphometric parameters, and quantify dot count, mean intensity, total area, intensity sum, and per-dot area/intensity per compartment.
cell culture, breast cancer, fluorescence, HER2

Custom App development
Perfectly tailored image analysis solutions for your research.
You have a specific research question that needs to be answered? We offer custom development of image analysis pipelines for specific tasks, be it detection of cellular phenotypes or quantification of tissue structures. After discussing your goals with one of our experts, you will get a ready-to-use App and be a step closer to an impactful publication.